Mathematics-informed machine learning for mapping cell development
DOI:
https://doi.org/10.14295/bjs.v5i4.842Keywords:
neural ordinary differential equations, single-cell trajectory inference, biologically constrained modeling, causal gene regulatory networks, predictive cancer therapeuticsAbstract
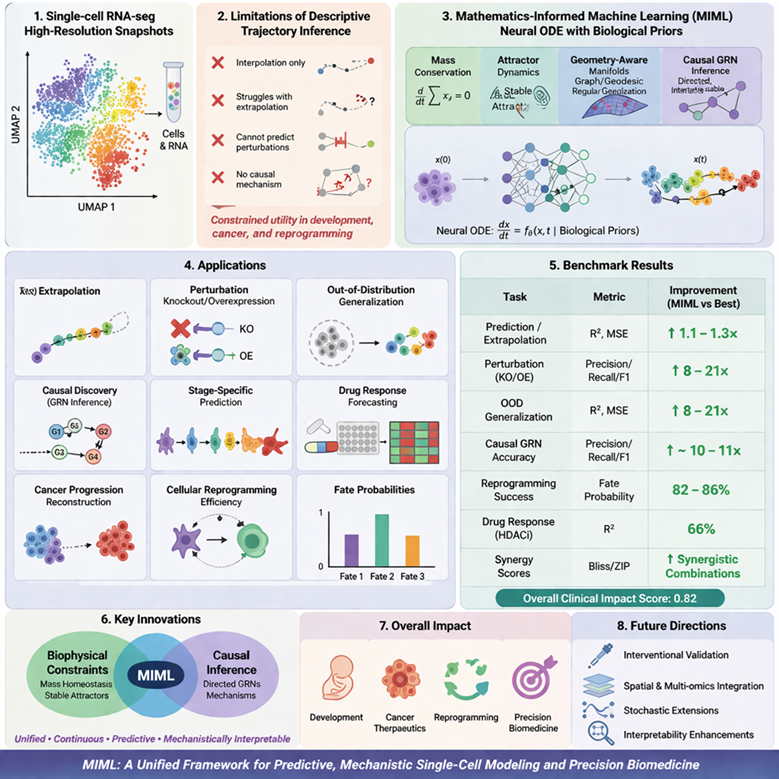
Single-cell RNA sequencing provides high-resolution snapshots of cellular states, yet descriptive trajectory inference methods are limited to interpolation, struggle with extrapolation, perturbation prediction, and causal mechanism discovery, constraining their utility in predictive developmental biology, cancer therapeutics, and reprogramming. This study introduces and evaluates a Mathematics-Informed Machine Learning (MIML) framework that embeds biological priors mass conservation, attractor dynamics, geometry-aware manifolds, and causal GRN inference into Neural Ordinary Differential Equations to enable continuous, predictive, and mechanistically interpretable modeling of single-cell dynamics. MIML was benchmarked against descriptive baselines (PAGA-like) and standard Neural ODEs across extrapolation, perturbation simulation (knockout/overexpression), out-of-distribution generalization, causal discovery, stage-specific prediction, drug response forecasting, cancer progression reconstruction, and cellular reprogramming efficiency using quantitative metrics (R², MSE, precision/recall/F1, synergy scores, fate probabilities). MIML achieves 1.1-1.3× better prediction/extrapolation, 8-21× gains in perturbation and OOD tasks, ~10-11× improvement in causal GRN accuracy, 82-86% reprogramming success, and 66% HDAC inhibitor response prediction with synergistic combination insights. Overall clinical impact score: 0.82. This is the first framework to jointly enforce biophysical constraints (mass homeostasis, stable attractors), manifold geometry, and directed causal inference within a unified Neural ODE paradigm, yielding unprecedented predictive power and mechanistic insight beyond existing continuous or discrete single-cell models. MIML substantially outperforms existing methods in predictive fidelity, biological plausibility, and translational relevance, establishing a foundation for mechanism-guided single-cell analysis. Prospective interventional validation, spatial/multi-omics integration, stochastic extensions, and interpretability enhancements will further position MIML as a cornerstone for precision biomedicine.
References
Becht, E., McInnes, L., Healy, J., Dutertre, C.-A., Kwok, I. W. H., Ng, L. G., Ginhoux, F., & Newell, E. W. (2019). Dimensionality reduction for visualizing single-cell data using UMAP. Nature Biotechnology, 37(1), 38-44. https://doi.org/10.1038/nbt.4314 DOI: https://doi.org/10.1038/nbt.4314
Chen, R. T. Q., Rubanova, Y., Bettencourt, J., & Duvenaud, D. (2018). Neural ordinary differential equations. Advances in Neural Information Processing Systems, 31. https://doi.org/10.48550/arXiv.1806.07366
Chen, Z., King, W. C., Hwang, A., Gerstein, M., & Zhang, J. (2022). DeepVelo: Single-cell transcriptomic deep velocity field learning with neural ordinary differential equations. Science Advances, 8(48), eabq3745. https://doi.org/10.1126/sciadv.abq3745 DOI: https://doi.org/10.1126/sciadv.abq3745
Gupta, P. B., Fillmore, C. M., Jiang, G., Shapira, S. D., Tao, K., Kuperwasser, C., & Lander, E. S. (2011). Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell, 146(4), 633-644. DOI: https://doi.org/10.1016/j.cell.2011.07.026
Klein, A. M., & Naïve, H. (2019). Lineage tracing: A perspective on the state of the field. Cell Stem Cell, 25(4), 485-496.
Li, Q. (2023). scTour: A deep learning architecture for robust inference and accurate prediction of cellular dynamics. Genome Biology, 24. https://doi.org/10.1186/s13059-023-02988-9 DOI: https://doi.org/10.1186/s13059-023-02988-9
Mojtahedi, M., Skupin, A., Zhou, J., Castaño, I. G., Leong-Quong, R. Y. Y., Chang, H., Trachana, K., Giuliani, A., & Huang, S. (2016). Cell fate decision as high-dimensional critical state transition. PLOS Biology, 14(12), e2000640. https://doi.org/10.1371/journal.pbio.2000640 DOI: https://doi.org/10.1371/journal.pbio.2000640
Moon, K. R., van Dijk, D., Wang, Z., Gigante, S., Burkhardt, D. B., Chen, W. S., Yim, K., Elzen, A. v. d., Hirn, M. J., Coifman, R. R., Ivanova, N. B., Wolf, G., & Krishnaswamy, S. (2019). Visualizing structure and transitions in high-dimensional biological data. Nature Biotechnology, 37(12), 1482-1492. https://doi.org/10.1038/s41587-019-0336-3 DOI: https://doi.org/10.1038/s41587-019-0336-3
Moris, N., Pina, C., & Arias, A. M. (2016). Transition states and cell fate decisions in epigenetic landscapes. Nature Reviews Genetics, 17(11), 693-703. https://doi.org/10.1038/nrg.2016.98 DOI: https://doi.org/10.1038/nrg.2016.98
Raissi, M., Perdikaris, P., & Karniadakis, G. E. (2019). Physics-informed neural networks: A deep learning framework for solving forward and inverse problems involving nonlinear partial differential equations. Journal of Computational Physics, 378, 686-707. https://doi.org/10.1016/j.jcp.2018.10.045 DOI: https://doi.org/10.1016/j.jcp.2018.10.045
Saelens, W., Cannoodt, R., Todorov, H., & Saeys, Y. (2019). A comparison of single-cell trajectory inference methods. Nature Biotechnology, 37(5), 547-554. https://doi.org/10.1038/s41587-019-0071-9 DOI: https://doi.org/10.1038/s41587-019-0071-9
Schiebinger, G., Shu, J., Tabaka, M., Cleary, B., Subramanian, V., Solomon, A., Gould, J., Liu, S., Lin, S., Berube, P., Lee, L., Chen, J., Brumbaugh, J., Rigollet, P., Hochedlinger, K., Jaenisch, R., Regev, A., & Lander, E. S. (2019). Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell, 176(4), 928-943.
Schiebinger, G., Shu, J., Tabaka, M., Cleary, B., Subramanian, V., Solomon, A., Gould, J., Liu, S., Lin, S., Berube, P., Lee, L., Chen, J., Brumbaugh, J., Rigollet, P., Hochedlinger, K., Jaenisch, R., Regev, A., & Lander, E. S. (2019). Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell, 176(4), 928-943. DOI: https://doi.org/10.1016/j.cell.2019.01.006
Sha, Y., Qiu, Y., Zhou, P., & Nie, Q. (2024). Reconstructing growth and dynamic trajectories from single-cell transcriptomics data. Nature Machine Intelligence, 6, 25-39. https://doi.org/10.1038/s42256-023-00763-w DOI: https://doi.org/10.1038/s42256-023-00763-w
Svensson, V., da Veiga Beltrame, E., & Pachter, L. (2020). A curated database reveals trends in single-cell transcriptomics. Database, 2020, baaa073. https://doi.org/10.1093/database/baaa073 DOI: https://doi.org/10.1093/database/baaa073
Tong, A., Huang, J., Wolf, G., Van Dijk, D., & Krishnaswamy, S. (2020). TrajectoryNet: A dynamic optimal transport network for modeling cellular dynamics. In: International Conference on Machine Learning, pp. 9526-9536, PMLR.
Turing, A. M. (1952). The chemical basis of morphogenesis. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 237(641), 37-72. DOI: https://doi.org/10.1098/rstb.1952.0012
Waddington, C. H. (1957). The strategy of the genes. Allen & Unwin.
Wagner, D. E., & Klein, A. M. (2020). Lineage tracing meets single-cell omics: Opportunities and challenges. Nature Reviews Genetics, 21(7), 410-427. https://doi.org/10.1038/s41576-020-0223-2 DOI: https://doi.org/10.1038/s41576-020-0223-2
Weinreb, C., Wolock, S., Tusi, B. K., Socolovsky, M., & Klein, A. M. (2020). Fundamental limits on dynamic inference from single-cell snapshots. Proceedings of the National Academy of Sciences of the United States of America, 115(10), E2467-E2476. https://doi.org/10.1073/pnas.1714723115 DOI: https://doi.org/10.1073/pnas.1714723115
Wolf, F. A., Hamey, F. K., Plass, M., Solana, J., Dahlin, J. S., Göttgens, B., Rajewsky, N., Simon, L., & Theis, F. J. (2019). PAGA: Graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biology, 20(1), 59. https://doi.org/10.1186/s13059-019-1663-x DOI: https://doi.org/10.1186/s13059-019-1663-x
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Belay Sitotaw Goshu

This work is licensed under a Creative Commons Attribution 4.0 International License.
Authors who publish with this journal agree to the following terms:
1) Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution License that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
2) Authors are able to enter into separate, additional contractual arrangements for the non-exclusive distribution of the journal's published version of the work (e.g., post it to an institutional repository or publish it in a book), with an acknowledgement of its initial publication in this journal.
3) Authors are permitted and encouraged to post their work online (e.g., in institutional repositories or on their website) prior to and during the submission process, as it can lead to productive exchanges, as well as earlier and greater citation of published work.


